Biodesign based on engineering principles
At the NUS Engineering Biology Lab, we focus on synthetic biology and apply engineering principles to design and build microbes with useful capabilities to address biomedical and sustainability challenges. We have special interest in engineering living biosensors for applications in health, sustainable biomanufacturing and DNA data storage. At the same time, we are developing foundational platform tools to accelerate the design and engineering of the microbes, making engineering biology more predictive and rational.
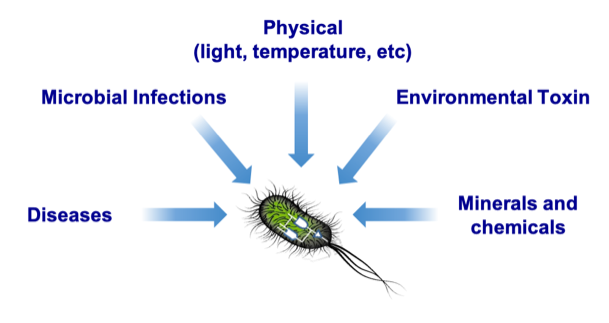
We are discovering and developing novel genetically encoded biosensors that, when incorporated into our engineered cells, allow them to react to different stimuli, e.g., sensors that respond to physical cues such as light/heat, sensors that detect molecules produced by pathogens, and sensors that detect chemicals. Layered with genetic circuits, we could then reprogram microbes to actuate different responses for different applications.
We engineer cell factories to produce several value-added proteins or chemicals (terpenoids, flavonoids etc.). In our lab, we have engineered the cells to produce bacterial collagen which has been reported to be a promising substitute to mammalian collagen and has been tested as non-toxic, biocompatible and with a cell-specific adhesion capability with improved spreading effects. We have also engineered cells to produce different flavonoids including vanillin which is main chemical compound of vanilla extract and can be used as chemical intermediate in the manufacture of several important drugs and other products.
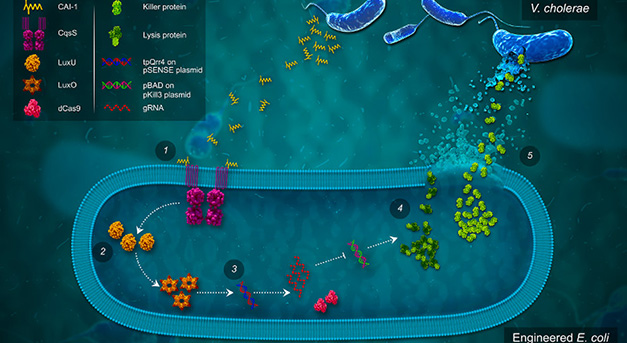
We have engineered microbes with the capability as diagnostic tool to sense and kill pathogens. Using a synthetic biology approach, we rationally design bacterial constructs to fight potentially deadly pathogens such as V. cholerae and engineer human commensal microbe to sense and kill antibiotic-resistant strain of P. aeruginosa. This strategy allows to exert an antibacterial effect only when it is needed, thus preventing development of resistance.
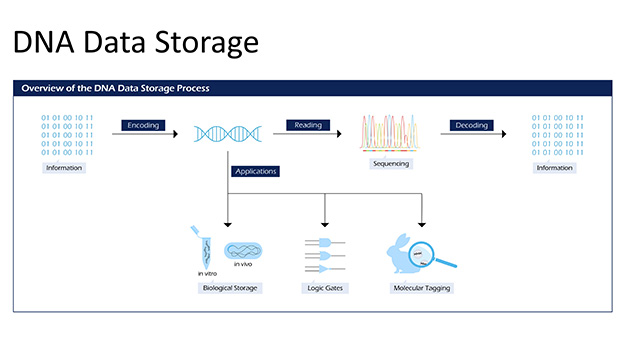
DNA data storage is an exciting new concept that aims to store our everyday information into the fabric of life itself – DNA. The immense density of DNA (215 petabytes per gram), its longevity (potentially millions of years in the appropriate conditions) as well as eternal biological relevance has led to significant interest in the area. We are developing synbio technology to exploit the use of DNA for data storage. The intent is to address the pressing issue with the impending shortage of silica necessary for manufacturing storage devices required to accommodate our projected data storage requirements. We in Poh lab are utilizing novel optogenetic tools, barcoding of bacterial cells, and spatial transcriptomics to provide an alternative approach to storing information into DNA – analogous to creating a camera for capturing and storing images directly into DNA itself.
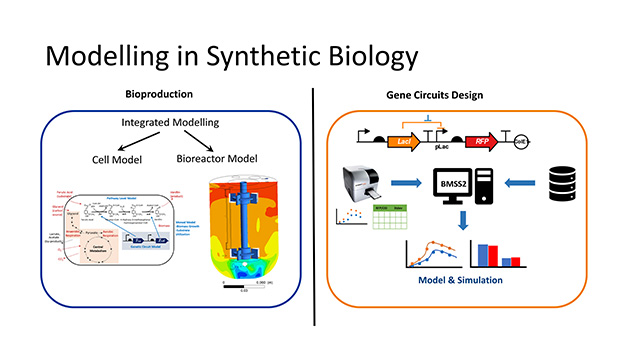
We develop mechanistic models and apply model-driven approach to enable the rational optimization of experimental designs and provide insights into the system behaviours. In our lab, mathematical modelling forms an integral part of most of the projects, e.g., biosensors and bioproduction in guiding the rational design of different constructs and providing quantitative insights into system performances. Modelling tools have also been developed to facilitate the model development and analysis.
The Engineering Biology Lab at NUS focuses on Synthetic Biology in which we apply engineering principles to design and build microbes with useful capabilities for biomedical and industrial applications, with special interest in engineering novel living biosensors for applications in health, sustainable biomanufacturing, and DNA data storage. To this end, we have been “reprogramming” microbes as living biosensors to fight infectious causing pathogen, light/thermal controllable living biosensors for biomanufacturing, and biosensors for ultra-high throughput screening.
At the same time, we are developing foundational platform tools to accelerate the design and engineering of the microbes along the design-build-test-learn cycle. This includes computer aided design and modelling tools for SynBio and biofoundry for rapid prototyping We combine both experimental and computational approaches in engineering our microbes.
We are highly motivated to make engineering of biology more efficient and predictive so that we can scale complexity in order to create novel solutions to tackle global challenges.